| |
Corresponding author: *; equal contribution: **; undergraduates: underlined.
Year 2026
72. Khan, G., Trinh, C.T.*, 2026, to be submitted.
71. Dooley, D., Trinh, C.T.*, 2026, to be submitted.
70. Dooley, D., Boyd, H., Trinh, C.T.*, 2026, to be submitted.
69. Dooley, D.**, Boyd, H.**, Trinh, C.T.*, 2026, to be submitted.
68. Cotter, C.J., Trinh, C.T.*, 2026, to be submitted.
67. Cotter, C.J., Carper, D.L., Giannone, R.J, Trinh, C.T.*, 2026, in review.
66. Dooley, D., Carper, D.L., Giannone, R.J., Trinh, C.T.*, 2026, Proteome Reallocation Reveals Dynamics and Mechanisms of Phage Infection and Defense in Staphylococcus aureus, bioRxiv. 
Year 2025
65. Khan, G.**, Sanford, C.**, Trinh, C.T.*, 2025, Medium Optimization to Produce Heterologous Proteins: Experimental Design, Metabolic Network Analysis, and Machine Learning, Biotechnol Advances, 86, 108738. 
64. Ha, K., Ryu, S., Trinh, C.T*, 2025. Alpha ketoacid decarboxylases: Diversity, structures, reaction mechanisms, and applications for biomanufacturing of platform chemicals and fuels, Biotechnol Advances, 81:108531. 
Year 2024
63. Dooley, D., Ryu, S., Jackson, E., Giannone, R., Dien, B., Trinh, C.T.*, 2024. Expanded Genome and Proteome Reallocation in A Novel, Robust Bacillus coagulans Capable of Utilizing Pentose and Hexose Sugars, mSystems,9(11): e00952-24. 
62. Cotter, C.J., Trinh, C.T.*, 2024. CRISPR-GRIT: Guide-RNAs with Integrated Repair Templates Enable Precise Multiplexed Genome Editing in the Diploid Fungal Pathogen Candida albicans, CRISPR Journal, 7(6):385-394. Cover Art. 
61. Klein, B.C., Scheidemantle, B., Hanes, R., Bartling, A., Grundl, N., Biddy, M., Tao, L., Trinh, C.T., Guss, A., Wyman, C.E., Ragauskas, A.J., Webb, E.G., Davison, B.H., Cai, C.M.*, 2024. Economics and global warming potential of a commercial-scale delignifying biorefinery based on Co-solvent Enhanced Lignocellulosic Fractionation, Energy Environ Sci, 17, 1202-1215. 
60. Seo, H., Castro, G., Trinh, C.T.*, 2024. Engineering a Syntrophic Escherichia coli Coculture for Compartmentalized de novo Biosynthesis of Isobutyl Butyrate from Mixed Sugars, ACS Synthetic Biology,13 (1), 259-268. 
Year 2023
59. Mendoza, B., Zheng, X., Clements J.§, Cotter, C., Trinh, C.T.*, 2023. Potency of CRISPR-Cas Antifungals Is Enhanced by Co-targeting DNA Repair and Growth Regulatory Machinery at the Genetic Level, ACS Infectious Diseases, 9: 2494–2503.
58. Walker, C., Mortensen, M., Poudel, B., Cotter, C., Okekeogbu, I., Ryu, S., Khomami, B., Giannone, R.J., Laursen, S., Trinh, C.T.*, 2023. Proteomes Reveal Metabolic Capabilities of Yarrowia lipolytica for Biological Upcycling of Polyethylene into High-Value Chemicals, mSystems, e00741-23. 
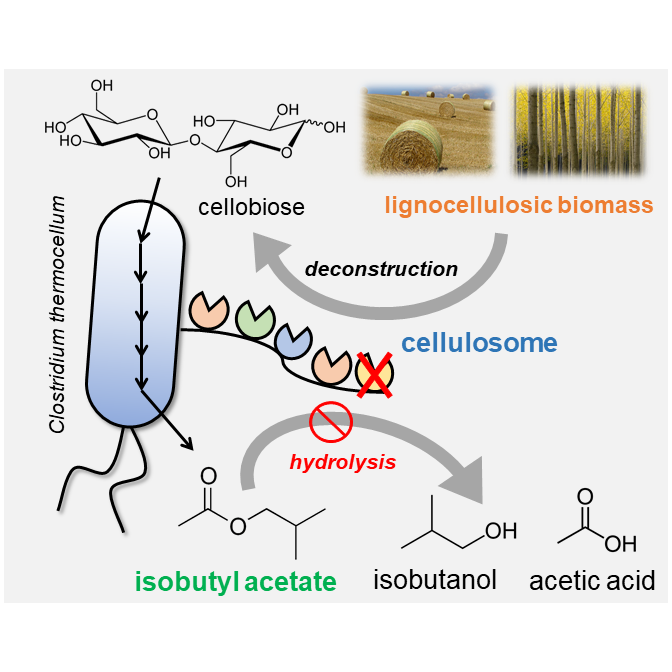
57. Seo, H., Singh, P., Wyman, C., Cai, C., Trinh, C.T.*, 2023. Rewiring metabolism of Clostridium thermocellum for consolidated bioprocessing of lignocellulosic biomass poplar to produce short-chain esters, Bioresource Technol, 384 (2023): 129263.
Year 2022
56. Walker, C., Ryu, S., Garica, S., Dooley, D., Mendoza, B., Trinh, C.T.* 2022. Gene Co-expression Connectivity Predicts Gene Targets Underlying High Ionic Liquid Tolerance in Yarrowia lipolytica, mSystems, 7(4): e00348-22 . .
55. Mendoza, B.**, Fry, T.**, Dooley, D.**, Herman J., Trinh, C.T.*, 2022. CASPER: An integrated software platform for rapid development of CRISPR tools, CRISPR J, 5(4): 609-617  . .
54. Seo, H., Giannone, R.J., Yang, Y.H., Trinh, C.T.* 2022. Proteome reallocation enables the selective de novo biosynthesis of non-linear, branched-chain acetate esters, Metab Eng, 73, 38-49.
53. Lynd, L.R., Beckham, G.T., Guss, A.M., Jayakody, L., Karp, E.M., Maranas, C., McCormick, R.L., Amador-Noguez, D., Bomble, Y., Davison, B.H., Foster, C., Himmel, M., Holwerda, E., Laser, M.S., Ng, C.Y., Olson, D.G., Román-Leshkov, Y., Trinh, C.T., Tuskan, G.A., Upadhayay, V., Vardon, D.R., Wang, L., Wyman, C.A., 2022. Toward low-cost biological and hybrid biological/catalytic conversion of cellulosic biomass to fuels, Energy Environ Sci, 15: 938-990. 
52. Lee, J., Trinh, C.T.*, 2022.Controlling Selectivity of Modular Microbial Biosynthesis of Butyryl-CoA-Derived Designer Esters, Metab Eng, 69: 262-274. 
Year 2021
51. Lee, J., Seo, H., Young, C., Trinh, C.T.*, 2021. Probing Specificities of Alcohol Acyltransferases for Designer Ester Biosynthesis with a High-Throughput Microbial Screening Platform, Biotechnol Bioeng, 1-13.
50. Walker, C., Dien, B., Giannone, R.J., Slininger, P., Thompson, S., Trinh, C.T.*, 2021. Exploring Proteomes of Robust Yarrowia lipolytica Isolates Cultivated in Biomass Hydrolysate Reveal Key Processes Impacting Mixed Sugar Utilization, Lipid Accumulation, and Degradation, mSystems, 6(4): e00443-21. 
49. Garcia, S., Trinh, C.T.*, 2021. Computational Design and Analysis of Modular Cells for Large Libraries of Exchangeable Product Synthesis Modules, Metab Eng, 67: 453-463. 
48. Poudel, S., Cope, A. L., O’Dell, K., Guss, A. M., Seo, H., Trinh, C. T., Hettich, R. L., 2021. An integrated guilt-by-association approach to characterize proteins of unknown function (PUFs) in Clostridium thermocellum DSM 1313 as potential genetic engineering targets, BBIO, 14 (116): 1-19. 
47. Seo, H., Lee, J., Dunlap, N., Giannone, R., Trinh, C.T.*, 2021. Engineering Promiscuity of Chloramphenicol Acetyltransferase for Microbial Designer Ester Biosynthesis, Metab Eng, 66: 179-190. 
Year 2020
46. Yang, X, Medford, J.I., Markel, K., Shih, P., De Paoli, H.C., Trinh, C.T., McCormick, A.J., Ployet, R., Hussey, S.G., Myburg, A.A., Jensen, P.E., Hassan, M.M., Zhang, J., Muchero, W., Kalluri, U.C., Yin, H., Zhuo, R., Abraham, P., Chen, J.G., Weston, D., Yang, Y.., Liu, D., Li, Y., Labbe, J., Yang, B., Lee, J., Cottingham, R.W., Martin, A., Lu, M., Tschaplinski, T.J., Y, G., Lu, H., Ranjan, P., Mitchell, J., Wullschleger, S.D., Tuskan, G.A., 2020. Plant Biosystems Design Research Roadmap 1.0, BioDesign Research, 2020, Article ID 8051764. 
45. Garcia, S., Thompson, R.A., Giannone, R., Dash, S., Maranas, C., Trinh, C.T.*, 2020. Development of a genome-scale metabolic model of Clostridium thermocellum and its applications for integration of multi-omics datasets and strain design, Frontiers Bioeng Biotechnol, 8: 1-16. 
44. Garcia, S., Trinh, C.T.*, 2020. Harnessing natural modularity of cellular metabolism to design a modular chassis cell for a diverse class of products by using goal attainment optimization, ACS Synthetic Biology, 9(7): 1665–1681. |
|